13 Sticking together
Sticking together
In this practical, you will practice building your own model of collective behaviour, based on the one you saw at the end of the lecture:
The example above is a implemented in Javascript, a programming language that is widely used for web development. It is easy to share with others, interactive, and surprisingly fast. But, it’s not the most “professional” programming language. Plus, at this stage of the course there is no point in learning yet another programming language, as you are here to learn about modeling biology. So we will stick to Python.
First, let’s discuss how we can let individuals walk around in space.
13.1 Steering
We can represent a moving individual in space as a point with a position and a velocity. The position is represented by two coordinates, \(x\) and \(y\), and the velocity is represented by two components, \(v_x\) and \(v_y\). All movement that this individual can do, will be a matter of repeatedly updating its position based on their velocity:
To model such a vector in python, we can simply define a base point with an x- and y-coordinate, and a velocity vector with an x- and y-component. The position of the individual can then be updated by adding the velocity to the position. Combining that with a function that draws an arrow in Python, we get the following code:
import numpy as np
import matplotlib.pyplot as plt
# Enable interactive mode for matplotlib
plt.ion()
# Setup figure and axis for plotting the arrow
fig, ax = plt.subplots(figsize=(8, 4))
ax.set_xlim(0, 600) # x-axis limits
ax.set_ylim(0, 250) # y-axis limits
ax.set_aspect('equal') # Keep aspect ratio square
ax.set_facecolor('#f0f0f0') # Background color
ax.set_title("A moving vector with an arrowhead") # Title
# Initial position and velocity
x, y = 250.0, 180.0 # Position coordinates
vx, vy = 5.0, 10.5 # Velocity components
def draw_arrow(x, y, vx, vy):
"""
Draws an arrow at position (x, y) with velocity (vx, vy).
"""
ax.clear()
ax.set_xlim(0, 600)
ax.set_ylim(0, 250)
ax.set_aspect('equal')
ax.set_facecolor('#f0f0f0')
ax.set_title("A moving vector with an arrowhead")
# Normalize velocity for drawing the arrow
dx = vx*5
dy = vy*5
# Arrow shaft
end_x = x + dx
end_y = y + dy
# Arrowhead calculation
angle = np.arctan2(dy, dx)
angle_offset = np.pi / 7
hx1_x = end_x - np.cos(angle - angle_offset)
hx1_y = end_y - np.sin(angle - angle_offset)
hx2_x = end_x - np.cos(angle + angle_offset)
hx2_y = end_y - np.sin(angle + angle_offset)
# Draw shaft
ax.quiver(x, y, dx, dy, angles='xy', scale_units='xy', scale=1, color='#007acc', width=0.005)
# Draw base point
ax.plot(x, y, 'o', color='#333')
# Labels
ax.text(x+10, y+10, f"x = {x:.2f}")
ax.text(x+10, y-5, f"y = {y:.2f}")
ax.text(end_x + 10, end_y - 20, f"vₓ = {vx:.2f}")
ax.text(end_x + 10, end_y, f"vᵧ = {vy:.2f}")
plt.draw()
plt.pause(0.03)
# Animation loop: update position by velocity
for i in range(500):
x += vx*0.1 # Update x position
y += vy*0.1 # Update y position
# Wrap around edges
x %= 600
y %= 250
draw_arrow(x, y, vx, vy)
plt.ioff()Exercise 13.1 (Playing with steering arrows - Mathematical thinking)
Copy-paste the code above, study it for a few minutes, and run it.
- What can you do to make the arrow accelerate?
To rotate a vector, we can use the following trigonometrical equations, where \(\theta\) is the angle of rotation:
\[ \begin{aligned} x_{new} = x \cdot \cos(\theta) - y \cdot \sin(\theta) \newline y_{new} = x \cdot \sin(\theta) + y \cdot \cos(\theta) \end{aligned} \]
Use the equation above to rotate the velocity vector in the code by a small angle every timestep. What happens?
Modeling 1 individual is not very exciting. Think about what the code above would look like if you had more than 1 individual. Discuss this with other students and/or Bram.
Moving “cells”
In this practical, you will practice with modeling individuals in space by modifying a Python code based on the foraging cells shown at the beginning. To accommodate for many cells, we will define a new Cell class, embedded in a Simulation class.1
First, read the code yourself (you can ignore the Visualisation class), and see if you can get it running on your own laptop.
###
# PRACTICAL 1 | "Every cell for themselves?"
# This is the starting code. Follow the instructions in the practical to complete the code.
# If you get stuck, you can look at the final code in `foraging_for_resources_final.py`, or ask
# Bram.
#
# The structure of this code is as follows:
# 1. Imports and parameters
# 2. Simulation class
# 3. Cell class
# 4. Visualisation class (you do not need to change this)
#
###
# 1. IMPORTS AND PARAMETERS
# Libraries
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.widgets import Slider
# Parameters for simulation
WORLD_SIZE = 200 # Width / height of the world (size of grid and possible coordinates for cells)
MAX_VELOCITY = 0.3 # Maximum velocity magnitude
MAX_FORCE = 0.3 # Maximum force magnitude
RANDOM_MOVEMENT = 0.01 # Random movement factor to add some noise to the cell's movement
# Parameters for display
DRAW_ARROW = True # Draw the arrows showing the velocity direction of the cells
INIT_CELLS = 20 # Initial number of cells in the simulation
DISPLAY_INTERVAL = 1 # Frequency with which the plot is updated (e.g., every 10 timesteps can speed things up)
# 1. MAIN LOOP (using functions and classes defined below)
def main():
"""Main function to set up and run the simulation."""
# NOTE: The `Visualisation` class is responsible for managing the visualization
# of the simulation, including creating plots, updating them, and handling
# user interactions like the slider. As this has nothing to do with modeling
# per se, understanding this code is not necessary, but it can be fun to look
# at if you are interested.
num_cells = INIT_CELLS
sim = Simulation(num_cells)
plt.ion()
vis = Visualisation(sim)
def update_cells(val):
sim.initialise_cells(int(vis.slider.val))
vis.redraw_plot(sim)
# Connect the slider to the update function
vis.slider.on_changed(update_cells)
# Run simulation
for t in range(1, 10000):
sim.simulate_step()
if(t % DISPLAY_INTERVAL == 0):
# As long as only cells move, update only positions and timestamp
vis.update_plot(sim)
vis.ax.set_title(f"Timestep: {t}")
vis.fig.canvas.draw_idle()
plt.pause(10e-20)
if(sim.redraw):
# When more has changes (e.g. number of cells or target position), redraw the plot
vis.redraw_plot(sim)
sim.redraw = False # Make sure it doesn't keep redrawing if not necessary
# Keep the final plot open
plt.ioff()
# plt.show()
# 2. SIMULATION CLASS
class Simulation:
"""Manages the grid, cells, target, and simulation logic."""
def __init__(self, num_cells):
# Initialise a grid for the simulation
self.grid = np.zeros((WORLD_SIZE, WORLD_SIZE)) # Initialise an empty grid
self.fill_grid(self.grid, 0, 0, 0, 0) # Fill grid with values (currently just 1s)
# Initialise a population of cells
self.cells = []
self.initialise_cells(num_cells)
# Place a 'target' in the middle
self.target_position = [WORLD_SIZE/2, WORLD_SIZE/2]
# A flag to only rebuild the plot when necessary (e.g. when the number of cells changes)
self.redraw = False
def simulate_step(self):
"""Simulate one timestep of the simulation."""
for cell in self.cells:
# Actions taken by each cell. Most of them are still undefined, so you can implement them yourself.
self.move_towards_dot(cell)
if self.check_target_reached(cell):
print(f"Target reached!")
self.reproduce_cell(cell)
self.redraw = True
#self.avoid_collision(cell)
#self.stick_to_close(cell)
#self.find_peak(cell)
# Apply forces and update position
cell.apply_forces()
cell.update_position()
# Limit velocity to the maximum allowed
cell.vx = np.clip(cell.vx, -MAX_VELOCITY, MAX_VELOCITY)
cell.vy = np.clip(cell.vy, -MAX_VELOCITY, MAX_VELOCITY)
def initialise_cells(self, num_cells):
"""Initialise the cells with random positions and velocities."""
self.cells = []
for _ in range(num_cells):
x = np.random.uniform(0, WORLD_SIZE)
y = np.random.uniform(0, WORLD_SIZE)
vx = np.random.uniform(-1, 1)
vy = np.random.uniform(-1, 1)
self.cells.append(Cell(x, y, vx, vy))
def fill_grid(self, grid, mean_x, mean_y, std_dev, noise=0):
"""
Write a function that takes the 2D grid and fills it with values representing
a Gaussian (normal) distribution centered at (mean_x, mean_y). See
if you can use the 'noise' argument to randomise the gaussian distribution a bit.
Hint: e^{-x^2} yields a bell curve centered around 0.
"""
for i in range(WORLD_SIZE):
for j in range(WORLD_SIZE):
x = i / (WORLD_SIZE - 1)
y = j / (WORLD_SIZE - 1)
grid[i, j] = 1 # This is 1 in the example, but should be a Gaussian distribution
# Normalize the grid to keep the total resource concentration the same
self.grid = grid
def find_peak(self, cell):
"""Make the cell move towards the peak of the resource gradient with a random walk."""
# Convert cell position to grid indices, as well as the previous position
grid_x = int(cell.x) % WORLD_SIZE
grid_y = int(cell.y) % WORLD_SIZE
next_x = (int(cell.x + 30*cell.vx) + WORLD_SIZE) % WORLD_SIZE
next_y = (int(cell.y + 30*cell.vy) + WORLD_SIZE) % WORLD_SIZE
def avoid_collision(self, cell):
"""Implement a simple collision avoidance mechanism. You can do so by
checking if this individual overlaps with another individual, and if so,
applying a repulsion force to the individual apposing the overlapping
direction."""
for other_cell in self.cells:
if other_cell is not cell:
# Calculate the distance between the two cells
dx = cell.x - other_cell.x
dy = cell.y - other_cell.y
distance = np.sqrt(dx**2 + dy**2)
def stick_to_close(self, cell):
"""Implement an attraction to cells that are nearby (but not overlapping)"""
for other_cell in self.cells:
if other_cell is not cell:
# Calculate the distance between the two cells
dx = cell.x - other_cell.x
dy = cell.y - other_cell.y
distance = np.sqrt(dx**2 + dy**2)
def move_towards_dot(self, cell):
"""
Write your own function that applies forces in the direction of the dot.
Try to think of a way to apply the same force to every cell irrespective
of the distance to the dot, such that the cells move towards the dot at
the same speed.
To get you started, the function already calculates dx and dy, which are
the distances to the target position in the x and y direction, respectively.
"""
# Calculate dx and dy
dx = self.target_position[0] - cell.x
dy = self.target_position[1] - cell.y
def check_target_reached(self, cell):
"""
Write your own function that checks if this cell has reached the target position.
You can do this by calculating the distance between the cell and the target.
If the distance is smaller than a certain threshold (e.g., 3 units), return True.
Otherwise, return False.
"""
return(False) # Dummy 'return' value.
def reproduce_cell(self, cell):
"""
Write your own function that reproduces this cell. Think
about what it should inherit, and what it should *not* inherit.
To keep the number of cell constant, you can first throw away a random cell.
"""
# Reproduce: Create a new cell with the same properties as the current cell
return(False) # Dummy 'return' value.
# 3. CELL CLASS
class Cell:
"""Represents an individual cell in the simulation."""
def __init__(self, x, y, vx, vy):
self.x = x
self.y = y
self.vx = vx
self.vy = vy
self.ax = 0
self.ay = 0
self.stickiness = 0.01 # Initial stickiness, can be adjusted later
def update_position(self):
"""Update the cell's position based on its velocity."""
self.x = (self.x + self.vx ) % WORLD_SIZE # Wrap around the world
self.y = (self.y + self.vy ) % WORLD_SIZE # Wrap around the world
def apply_forces(self):
"""Apply a force to the cell, updating its velocity."""
self.ax = np.clip(self.ax, -MAX_FORCE, MAX_FORCE)
self.ay = np.clip(self.ay, -MAX_FORCE, MAX_FORCE)
self.vx += self.ax + RANDOM_MOVEMENT * np.random.uniform(-1, 1)
self.vy += self.ay + RANDOM_MOVEMENT * np.random.uniform(-1, 1)
# Apply drag to slow down the cell naturally
self.ax = 0
self.ay = 0
# Visualisation class for showing the individuals and the grid. For the practical, you do not need to change this.
class Visualisation:
def __init__(self, sim):
fig, ax = plt.subplots(figsize=(6, 6))
self.cell_x = [cell.x for cell in sim.cells]
self.cell_y = [cell.y for cell in sim.cells]
self.cell_vx = np.array([cell.vx for cell in sim.cells])
self.cell_vy = np.array([cell.vy for cell in sim.cells])
self.cell_stickiness = np.array([cell.stickiness for cell in sim.cells])
# Colour cells by stickiness using inferno colormap
self.cell_scatter = ax.scatter(self.cell_x, self.cell_y, c=self.cell_stickiness, cmap='inferno', s=50, edgecolor='white', vmin=0, vmax=1)
if(DRAW_ARROW): self.cell_quiver = ax.quiver(self.cell_x, self.cell_y, self.cell_vx * 0.5, self.cell_vy * 0.5, angles='xy', scale_units='xy', scale=0.02, color='white')
plt.subplots_adjust(bottom=0.2)
ax.set_xlim(0, WORLD_SIZE)
ax.set_ylim(0, WORLD_SIZE)
ax.set_aspect('equal', adjustable='box')
ax.set_title(f"Timestep: 0")
ax.set_xlabel("X")
ax.set_ylabel("Y")
target_point=ax.scatter(sim.target_position[0], sim.target_position[1], c='purple', s=100, edgecolor='red')
grid_im=ax.imshow(sim.grid.T, extent=(0, WORLD_SIZE, 0, WORLD_SIZE), origin='lower', cmap='viridis', alpha=1.0)
self.fig = fig
self.ax = ax
self.target_point = target_point
self.grid_im = grid_im
# Add a slider for selecting the number of cells
ax_slider = plt.axes([0.2, 0.05, 0.6, 0.03])
self.slider = Slider(ax_slider, 'Cells', 1, 1000, valinit=len(sim.cells), valstep=1)
def update_cell_positions(self, sim):
"""Update the positions of the cells in the visualisation."""
self.cell_x = [cell.x for cell in sim.cells]
self.cell_y = [cell.y for cell in sim.cells]
self.cell_vx = np.array([cell.vx for cell in sim.cells])
self.cell_vy = np.array([cell.vy for cell in sim.cells])
self.cell_stickiness = np.array([cell.stickiness for cell in sim.cells])
def update_plot(self, sim):
self.update_cell_positions(sim)
self.cell_scatter.set_offsets(np.c_[self.cell_x,self.cell_y])
self.cell_scatter.set_array(self.cell_stickiness)
if(DRAW_ARROW):
self.cell_quiver.set_offsets(np.c_[self.cell_x, self.cell_y])
self.cell_quiver.set_UVC(self.cell_vx * 0.5, self.cell_vy * 0.5)
def redraw_plot(self, sim):
self.update_cell_positions(sim)
cell_scatter_new = self.ax.scatter(self.cell_x, self.cell_y, c=self.cell_stickiness, cmap='inferno', s=50, edgecolor='white', vmin=0, vmax=1)
if(DRAW_ARROW):
cell_quiver_new = self.ax.quiver(self.cell_x, self.cell_y, self.cell_vx * 0.15, self.cell_vy * 0.15, angles='xy', scale_units='xy', scale=0.02, color='white')
self.cell_quiver.remove()
self.cell_quiver = cell_quiver_new
self.cell_scatter.remove()
self.fig.canvas.draw_idle()
self.cell_scatter = cell_scatter_new
self.grid_im.remove()
self.grid_im = self.ax.imshow(sim.grid.T, extent=(0, WORLD_SIZE, 0, WORLD_SIZE), origin='lower', cmap='viridis', alpha=1.0)
self.target_point.remove()
self.target_point=self.ax.scatter(sim.target_position[0], sim.target_position[1], c='purple', s=100, edgecolor='red')
plt.pause(10e-20)
# 4. Execute the main loop
if __name__ == "__main__":
# with cProfile.Profile() as pr:
main()
# pr.print_stats()
Make sure you inspect the code. What features does the Simulation class have? What features does a Cell have?
As you can see if you inspected the code properly, many functions are left empty (or at least do not do anything yet). You will start filling these with your own code.
13.2 Moving the target
If you run the code, you will see a purple dot (with a red outline). This may represent a “target”. It could represent a resource patch for bacteria, but it could also be a piece of fruit for a monkey (at this point, the model is still very abstract, so both could be true). Let’s make the target change position around after an individual touches it.
Exercise 13.2 (Playing with steering arrows - Algorithmic/mathematical thinking) To help you on your way, first answer the following questions for yourself:
- How can you calculate the distance between an individual and the orange dot?
- When is an individual close enough to the orange dot?
- How can we assign a random position to the dot?
A working code to steer the individuals towards the target is shown below. Note that the acceleration that is applied is only small, otherwise the individuals will move in a straight line towards the target very rapidly.
First try it yourself. If you get stuck, ask Bram for help.
13.3 Reproduction
Let’s reward the individual that found the target. To do this, we can call the ‘Cell’ constructor to make a new cell, and add it to the list of cells:
new_cell = Cell(new_x, new_y, new_vx, new_vy, new_speed)
self.cells.append(new_cell)Exercise 13.3 (The birth of an arrow - Biological / algorithmic thinking) Consider which properties of the parent cell get inherited to the child:
- Should the exact position be inherited to offspring? (yes/no)
- Should the offspring be placed nearby its parent? (yes/no)
- Does the velocity get inherited?
Note that depending on the scenario, the above questions may change. When a planktonic algea reproduces the daughter cells may inherit the velocity of the mother cell, but if a monkey gives birth, it does not make a lot of sense to talk about the ‘velocity’ of the mother. If we consider plants, we should not even consider velocity of the individuals at any stage of their life.
13.4 Collision detection
As the cells move towards the dot, you may notice that cells start overlapping quite a bit. Let’s implement a simple form of collision detection, where overlap is resolved by pushing cells away from each other. Answer the following questions to get on your way:
Exercise 13.4 (Overlapping circles - Algorithmic/mathematical thinking)
- How can you calculate the distance between two cells?
- When are two points overlapping?
- What can we do when two points overlap?
Now that we have implemented both target-finding, reproduction, and collision, we can study these individual mechanisms by commenting one or the other out. This is an important process in understanding a model. Try it for yourself!
13.5 Implementing a resource peak
As you may have noticed when reading the code, it also includes a grid. However, you don’t see this grid yet, as the function fill_grid currently sets every point to 1.
We can loop over the grid coordinates by using a double for-loop:
for i in range(WORLD_SIZE):
for j in range(WORLD_SIZE):
grid[i,j] = 1The above function loops over all the grid points, and set the value of each grid point to 1. Let’s use the function np.exp to calculate a Gaussian that we will place in the center of the grid. The fill_grid already takes as arguments the grid, a relative x-coordinate (0-1), a relative y-coordinate (0-1), and a standard deviation (0-1). Get started by answering the question below.
Exercise 13.5 (Bell curves in space - Mathematical thinking)
- The function \(e^{-x^2}\) gives a bell-curve centered around zero. How can you make it centered around a different value?
- Combined with the numpy function
np.exp, how can we use the equation in question a to create a Gaussian that is at the center of the grid?
13.6 Run and tumble
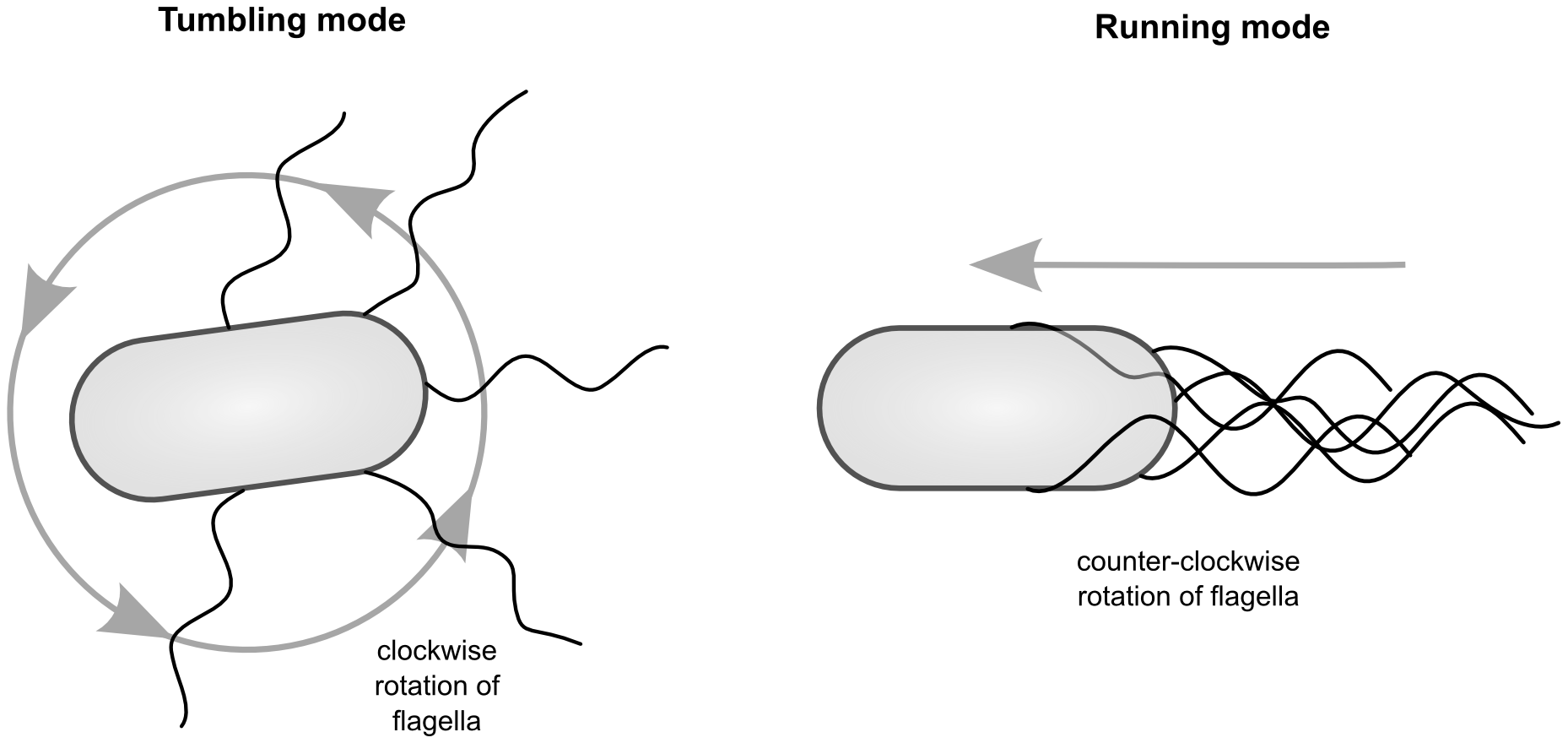
Bacterial cells are so small, that they cannot detect a gradient directly (in other words, they don’t know in which direction resources are higher!). Instead, bacteria often use a “run and tumble” strategy. When they are currently not detecting an increase in the concentration (over time), they tumble. If they do detect an increase, they keep moving in the same direction. This is a very simple strategy, but it can be very effective mechanism for chemotaxis.
In our earlier code of a single, moving vector, we rotated the arrow by changing the ‘angle’ variable. However, these cells do not have an angle parameters, but only a velocity vector with components \(v_x\) and \(v_y\). If we want to rotate the velocity vector, we can use the following equation (rooted in basic trigonometry):
\[ v_x' = v_x \cdot cos(\theta) - v_y \cdot sin(\theta) \\ v_y' = v_x \cdot sin(\theta) + v_y \cdot cos(\theta) \]
Where \(\theta\) is the angle we want the vector to rotate (in radians, not degrees!), and \(v_x\) and \(v_y\) are the components of the vector. With this in mind, let’s try and model chemotaxis.
Exercise 13.6 (Run and tumble - Biology / algorithmic thinking)
- Determine the concentration at the position of the cell, AND the predicted position of the cell after a small timestep (hint: use \(v_x\) and \(v_y\) to predict the future position! ask Bram if you get stuck)
- Make sure the future position is not outside of the grid! (hint: use Google, ChatGPT, or Copilot and figure out how the “modulo” operator works)
- If the future position has a higher concentration than the current position, keep moving in more or less the same direction, with a very small change.
- If the future position is a lower concentration, rotate the velocity vector a lot.
Study if your individuals can find the resource peak. Notice that depending on your implementation, it may or may not work. Make sure to carefully investigate why it does or does not work.
13.7 Sticking together
Cells sticking together can be implemented in multiple ways. Cells could be connected by a Newtonian spring, or we could simple make sure that cells that are close to each other are attracted to one another. In this case, we will use the latter method. Note that this is not very different from collision avoidance, but it is the other way around. In fact, we now have two opposing forces: cells are attracted to one another but do not want to overlap. This can be a bit finicky to get right, so feel free to explore. Ask for help if you get stuck.
13.8 1000 cells?
Try to run the model with 1000 cells. Also go back to the starting code again (without all the additions), and run this code with 1000 or cells too.
Exercise 13.7 (Algorithmic / computational thinking)
- What happens? Can you explain this?
This is far as this introduction to IBMs in python goes. Simple IBMs can be efficiently implemented in basic Python, but for more complex models, it is better to i) use numpy operations to speed up your Python code, or ii) use a faster programming language like C, Rust, or Javascript. For the last part of this pratical, we will study the Javascript version of this model. But before that, here is the final code that I ended up with:
13.9 Final Python code (for interested students)
This combination of individuals moving in continuous space, combined with a grid (e.g. with resources, or other environmental states) is a very useful way to make a spatially structured model.
###
# PRACTICAL 1 | "Every cell for themselves?"
# Things in this model that you have tried to implement yourself:
# 1. Implement collision avoidance
# 2. Implement reproduction
# 3. Implement a Gaussian grid
# 4. Implement "run and tumble"
# 5. Add noise to Gaussian, what happens?
# 5. Modify collision into STICKING (a little finicky)
# 6. Try it out with 500 cells...
###
###
# PRACTICAL 1 | PLENARY DISCUSSION
# What else was discussed in the plenary?
# 1. Why are grids so popular in modeling?
# 2. Tessellation of space
# 3. Automatic tessellation of space: quad tree
# 4. In the full model (javascript/Cacatoo), a quad tree is present, impacting performance
###
# 1. IMPORTS AND PARAMETERS
# Libraries
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.widgets import Slider
# Parameters for simulation
WORLD_SIZE = 200 # Width / height of the world (size of grid and possible coordinates for cells)
MAX_VELOCITY = 0.3 # Maximum velocity magnitude
MAX_FORCE = 0.3 # Maximum force magnitude
DRAG_COEFFICIENT = 0.01 # Friction to slow down the cell naturally
RANDOM_MOVEMENT = 0.01 # Random movement factor to add some noise to the cell's movement
CELL_STICKINESS_LOW = 0.0 # Minimal stickiness of cells in population
CELL_STICKINESS_HIGH = 0.10 # Maximal stickiness of cells in population
# Parameters for display
DRAW_ARROW = False # Draw the arrows showing the velocity direction of the cells
NOISE = 2 # Noise factor for the Gaussian grid (noise amount is raised to the power of this value)
INIT_CELLS = 64 # Initial number of cells in the simulation
SEASON_DURATION = 1000 # Duration of a season, after which the Gaussian grid is regenerated
DISPLAY_INTERVAL = 5
# 1. MAIN LOOP (using functions and classes defined below)
def main():
"""Main function to set up and run the simulation."""
# Initialise simulation and its # The `visualis` variable in the code snippet provided is actually
# a misspelling of the correct variable name `vis`, which stands
# for the `Visualisation` class instance. The `Visualisation`
# class is responsible for managing the visualization of the
# simulation, including creating plots, updating them, and
# handling user interactions like the slider.
num_cells = INIT_CELLS
sim = Simulation(num_cells)
plt.ion()
vis = Visualisation(sim)
def update_cells(val):
sim.initialise_cells(int(vis.slider.val))
vis.redraw_plot(sim)
# Connect the slider to the update function
vis.slider.on_changed(update_cells)
# Run simulation
for t in range(1, 10000):
sim.simulate_step()
if(t % DISPLAY_INTERVAL == 0):
vis.update_plot(sim)
vis.ax.set_title(f"Timestep: {t}")
vis.fig.canvas.draw_idle()
plt.pause(10e-20)
if(t % SEASON_DURATION==0):
sim.fill_grid(sim.grid, 0.2+np.random.uniform(0,0.6), 0.2+np.random.uniform(0,0.6), 0.1, NOISE)
vis.redraw_plot(sim)# Create Gaussian grid
# Update title and redraw the plot
# Keep the final plot open
plt.ioff()
# plt.show()
# 2. SIMULATION CLASS
class Simulation:
"""Manages the grid, cells, target, and simulation logic."""
def __init__(self, num_cells):
self.grid = np.zeros((WORLD_SIZE, WORLD_SIZE)) # Initialise an empty grid
self.cells = []
self.target_position = [WORLD_SIZE/3, WORLD_SIZE/3] # Initial target position at the center
self.target_position = [-1,-1]
self.fill_grid(self.grid, 0.5, 0.5, 0.1, NOISE) # Create Gaussian grid
self.initialise_cells(num_cells)
def simulate_step(self):
"""Simulate one timestep of the simulation."""
for cell in self.cells:
# Actions taken by each cell. Most of them are still undefined, so you can implement them yourself.
#self.move_towards_dot(cell)
#if self.check_target_reached(cell):
# print(f"Target reached! New target position: {self.target_position}")
# self.reproduce_cell(cell)
self.avoid_collision(cell)
self.stick_to_close(cell)
self.find_peak(cell)
# Apply drag force to acceleration
cell.ax += -DRAG_COEFFICIENT * cell.vx
cell.ay += -DRAG_COEFFICIENT * cell.vy
# Apply forces and update position
cell.apply_forces()
cell.update_position()
# Limit velocity to the maximum allowed
cell.vx = np.clip(cell.vx, -MAX_VELOCITY, MAX_VELOCITY)
cell.vy = np.clip(cell.vy, -MAX_VELOCITY, MAX_VELOCITY)
def initialise_cells(self, num_cells):
"""Initialise the cells with random positions and velocities."""
self.cells = []
for _ in range(num_cells):
x = np.random.uniform(0, WORLD_SIZE)
y = np.random.uniform(0, WORLD_SIZE)
vx = np.random.uniform(-1, 1)
vy = np.random.uniform(-1, 1)
self.cells.append(Cell(x, y, vx, vy))
def fill_grid(self, grid, mean_x, mean_y, std_dev, noise=0):
"""Creates a Gaussian distribution with noise on the grid."""
for i in range(WORLD_SIZE):
for j in range(WORLD_SIZE):
x = i / (WORLD_SIZE - 1)
y = j / (WORLD_SIZE - 1)
distance_squared = (x - mean_x)**2 + (y - mean_y)**2
grid[i, j] = np.exp(-distance_squared / (2 * std_dev**2)) * np.random.uniform(0.0, 1.0)**noise
# Normalize the grid to keep the total resource concentration the same
grid /= np.sum(grid)
self.grid = grid
def find_peak(self, cell):
"""Make the cell move towards the peak of the resource gradient with a random walk."""
# Convert cell position to grid indices, as well as the previous position
grid_x = int(cell.x) % WORLD_SIZE
grid_y = int(cell.y) % WORLD_SIZE
next_x = (int(cell.x + 10*cell.vx) + WORLD_SIZE) % WORLD_SIZE
next_y = (int(cell.y + 10*cell.vy) + WORLD_SIZE) % WORLD_SIZE
# Get the resource value at the cell's position, as well as the previous position
resource_value = self.grid[grid_x, grid_y]
resource_next = self.grid[next_x, next_y]
# Check if the cell is moving in the right direction
if resource_next > resource_value:
# Moving in the right direction: small random adjustment
angle = np.random.uniform(-0.1, 0.1) # Small angle change
else:
# Moving in the wrong direction: large random adjustment
angle = np.random.uniform(-np.pi*1.0, np.pi*1.0) # Large angle change
# Rotate the velocity vector by the random angle according to trigonometric rotation formulas
new_vx = cell.vx * np.cos(angle) - cell.vy * np.sin(angle)
new_vy = cell.vx * np.sin(angle) + cell.vy * np.cos(angle)
# Update the acceleration with the new velocity vector, such that the cell moves towards the peak
cell.vx = new_vx
cell.vy = new_vy
cell.ax += cell.vx
cell.ay += cell.vy
def avoid_collision(self, cell):
"""Avoidance forces to prevent cells from colliding."""
for other_cell in self.cells:
if other_cell is not cell:
# Calculate the distance between the two cells
dx = cell.x - other_cell.x
dy = cell.y - other_cell.y
distance = np.sqrt(dx**2 + dy**2)
# If the cells are too close, apply a repulsion force
if distance < 5.0 and distance > 0: # Threshold for "too close"
# Calculate the repulsion force proportional to the inverse of the distance
force_magnitude = (5.0 - distance) / distance
cell.ax += force_magnitude * dx * 100
cell.ay += force_magnitude * dy * 100
def stick_to_close(self, cell):
"""Stick to closeby cells."""
for other_cell in self.cells:
if other_cell is not cell:
# Calculate the distance between the two cells
dx = cell.x - other_cell.x
dy = cell.y - other_cell.y
distance = np.sqrt(dx**2 + dy**2)
# If the cells are too close, apply a repulsion force
if distance < 12 and distance > 5: # Threshold for "close"
# Calculate the repulsion force proportional to the inverse of the distance
cell.ax -= cell.stickiness * dx *10
cell.ay -= cell.stickiness * dy *10
def move_towards_dot(self, cell):
"""Apply forces in the direction of the dot."""
# Calculate dx and dy
dx = self.target_position[0] - cell.x
dy = self.target_position[1] - cell.y
# Calculate the distance to the target (pythagorean theorem)
distance = np.sqrt(dx**2 + dy**2)
# Normalize dx and dy
dx /= distance
dy /= distance
# Apply a small force towards the target
cell.ax += dx * 0.01
cell.ay += dy * 0.01
def check_target_reached(self, cell):
distance_to_target = np.sqrt((cell.x - self.target_position[0])**2 +
(cell.y - self.target_position[1])**2)
if distance_to_target < 3:
# Set a new target position
self.target_position = [np.random.uniform(0, WORLD_SIZE), np.random.uniform(0, WORLD_SIZE)]
return(True)
return(False)
def reproduce_cell(self, cell):
# Reproduce: Create a new cell with the same properties as the current cell
angle = np.random.uniform(0, 2 * np.pi)
radius = np.random.uniform(0.05, 1.5)
new_x = cell.x + radius * np.cos(angle)
new_y = cell.y + radius * np.sin(angle)
new_cell = Cell(new_x, new_y, cell.vx, cell.vy)
random_cell = np.random.choice(self.cells)
self.cells.remove(random_cell)
self.cells.append(new_cell)
# 3. CELL CLASS
class Cell:
"""Represents an individual cell in the simulation."""
def __init__(self, x, y, vx, vy):
self.x = x
self.y = y
self.vx = vx
self.vy = vy
self.ax = 0
self.ay = 0
if(np.random.uniform(0,1) < 0.5):
self.stickiness = CELL_STICKINESS_LOW
else:
self.stickiness = CELL_STICKINESS_HIGH
def update_position(self):
"""Update the cell's position based on its velocity."""
self.x = (self.x + self.vx ) % WORLD_SIZE # Wrap around the world
self.y = (self.y + self.vy ) % WORLD_SIZE # Wrap around the world
def apply_forces(self):
"""Apply a force to the cell, updating its velocity."""
self.ax = np.clip(self.ax, -MAX_FORCE, MAX_FORCE)
self.ay = np.clip(self.ay, -MAX_FORCE, MAX_FORCE)
self.vx += self.ax + RANDOM_MOVEMENT * np.random.uniform(-1, 1)
self.vy += self.ay + RANDOM_MOVEMENT * np.random.uniform(-1, 1)
self.ax = 0
self.ay = 0
# Visualisation class for showing the individuals and the grid. For the practical, you do not need to change this.
class Visualisation:
def __init__(self, sim):
fig, ax = plt.subplots(figsize=(6, 6))
self.cell_x = [cell.x for cell in sim.cells]
self.cell_y = [cell.y for cell in sim.cells]
self.cell_vx = np.array([cell.vx for cell in sim.cells])
self.cell_vy = np.array([cell.vy for cell in sim.cells])
self.cell_stickiness = np.array([cell.stickiness for cell in sim.cells])
# Colour cells by stickiness using inferno colormap
self.cell_scatter = ax.scatter(self.cell_x, self.cell_y, c=self.cell_stickiness, cmap='inferno', s=50, edgecolor='white', vmin=0, vmax=CELL_STICKINESS_HIGH*1.2)
if(DRAW_ARROW): self.cell_quiver = ax.quiver(self.cell_x, self.cell_y, self.cell_vx * 0.15, self.cell_vy * 0.15, angles='xy', scale_units='xy', scale=0.02, color='darkblue')
plt.subplots_adjust(bottom=0.2)
ax.set_xlim(0, WORLD_SIZE)
ax.set_ylim(0, WORLD_SIZE)
ax.set_aspect('equal', adjustable='box')
ax.set_title(f"Timestep: 0")
ax.set_xlabel("X")
ax.set_ylabel("Y")
target_point=ax.scatter(sim.target_position[0], sim.target_position[1], c='orange', s=50, edgecolor='white')
grid_im=ax.imshow(sim.grid.T, extent=(0, WORLD_SIZE, 0, WORLD_SIZE), origin='lower', cmap='viridis', alpha=1.0)
self.fig = fig
self.ax = ax
self.target_point = target_point
self.grid_im = grid_im
# Add a slider for selecting the number of cells
ax_slider = plt.axes([0.2, 0.05, 0.6, 0.03])
self.slider = Slider(ax_slider, 'Cells', 1, 1000, valinit=len(sim.cells), valstep=1)
def update_cell_positions(self, sim):
"""Update the positions of the cells in the visualisation."""
self.cell_x = [cell.x for cell in sim.cells]
self.cell_y = [cell.y for cell in sim.cells]
self.cell_vx = np.array([cell.vx for cell in sim.cells])
self.cell_vy = np.array([cell.vy for cell in sim.cells])
self.cell_stickiness = np.array([cell.stickiness for cell in sim.cells])
def update_plot(self, sim):
self.update_cell_positions(sim)
self.cell_scatter.set_offsets(np.c_[self.cell_x,self.cell_y])
self.cell_scatter.set_array(self.cell_stickiness)
if(DRAW_ARROW):
self.cell_quiver.set_offsets(np.c_[self.cell_x, self.cell_y])
self.cell_quiver.set_UVC(self.cell_vx * 0.15, self.cell_vy * 0.15)
def redraw_plot(self, sim):
self.update_cell_positions(sim)
cell_scatter_new = self.ax.scatter(self.cell_x, self.cell_y, c=self.cell_stickiness, cmap='inferno', s=50, edgecolor='white', vmin=0, vmax=CELL_STICKINESS_HIGH*1.2)
if(DRAW_ARROW):
cell_quiver_new = self.ax.quiver(self.cell_x, self.cell_y, self.cell_vx * 0.15, self.cell_vy * 0.15, angles='xy', scale_units='xy', scale=0.02, color='darkblue')
self.cell_quiver.remove()
self.cell_quiver = cell_quiver_new
self.cell_scatter.remove()
self.fig.canvas.draw_idle()
self.cell_scatter = cell_scatter_new
self.grid_im.remove()
self.grid_im = self.ax.imshow(sim.grid.T, extent=(0, WORLD_SIZE, 0, WORLD_SIZE), origin='lower', cmap='viridis', alpha=1.0)
plt.pause(0.01)
# 4. Execute the main loop
if __name__ == "__main__":
# with cProfile.Profile() as pr:
main()
# pr.print_stats()
13.10 Exploring the full Javascript model with evolution
The full model is implemented in Javascript, and can be found here.
In this full model, cells also reproduce every once in a while (when a “season” ends). Their reproductive success is shaped by the amount of resources at that position. Every time a cell reproduces there it inherits the parents stickiness, but it can also change this value a bit. This way, stickiness is an “evolvable” property on which natural selection will act. How much each cell is attracted to nearby cells depends on an internal “stickiness” parameter. Let the simulation run for some time.
Exercise 13.8 (The evolution of stickiness - Biology)
- What happens to the evolution of stickiness?
- Identify multiple advantages and disadvantages of stickiness.
- Given your answer in b., can you name an important parameter that may determine the balance bewteen the advantages and disadvantages of stickiness? See if you can test it using the options provided.
The code also contains a
Visualisationclass that uses thematplotliblibrary to draw the cells and their movement, which we have tuned to speed things up a bit. You do not need to understand this part of the code, but if you are interested feel free to check it out.↩︎