9 Gene regulation in space
Now we have some hope of making progress – Niehls Bohr
9.1 Auxin does everything
In this practical you will study the effects of auxin, a plant hormone, on gene regulation in the root. Auxin is a hormone that is virtually involved in all developmental processes in plants, from how the plant embryo is patterned, how the organs in flowers are determined, the spatial disposition of leaves, among many many others. In each context, auxin triggers specific responses, posing the question of how is it that the same molecule can trigger so many different responses. This specificity of auxin responses can be explained by the underlying regulatory networks. The auxin signalling pathway activates the auxin response factors, ARFs, which are transcription factors that regulate gene expression in response to auxin. Arabidopsis has 23 genes encoding for ARFs, some regulate gene expression positively and others negatively, and they are expressed in different tissues. It is this diversity in transcription factors activity which in part explains how auxin can regulate so many different things.
9.2 Auxin does opposite things in the root
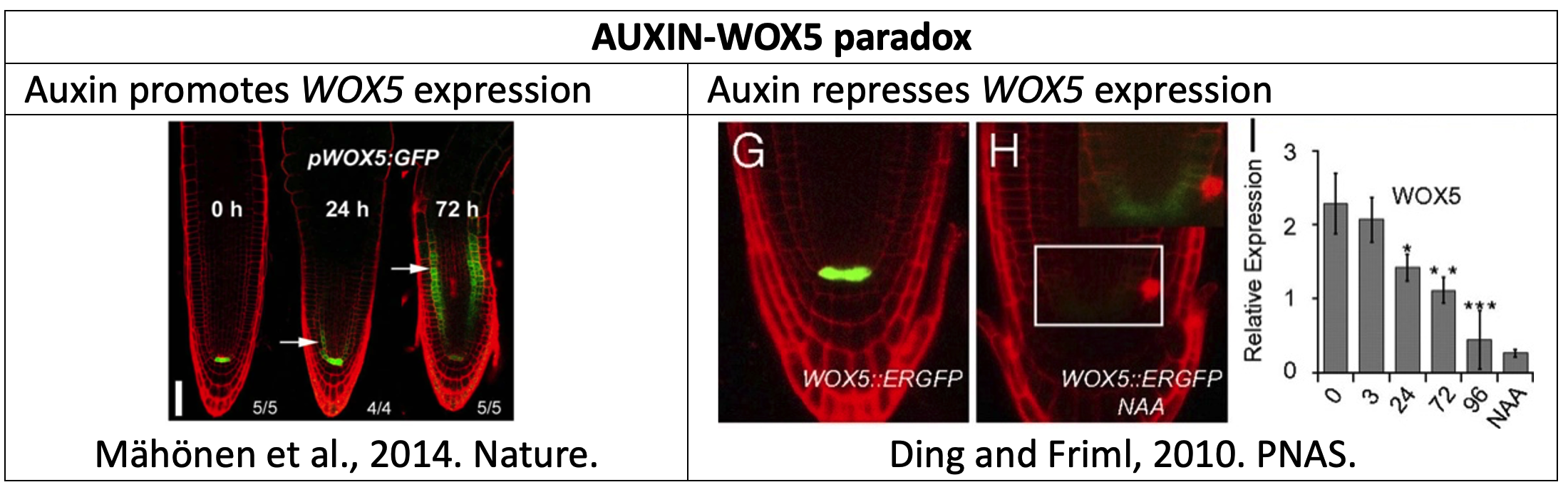
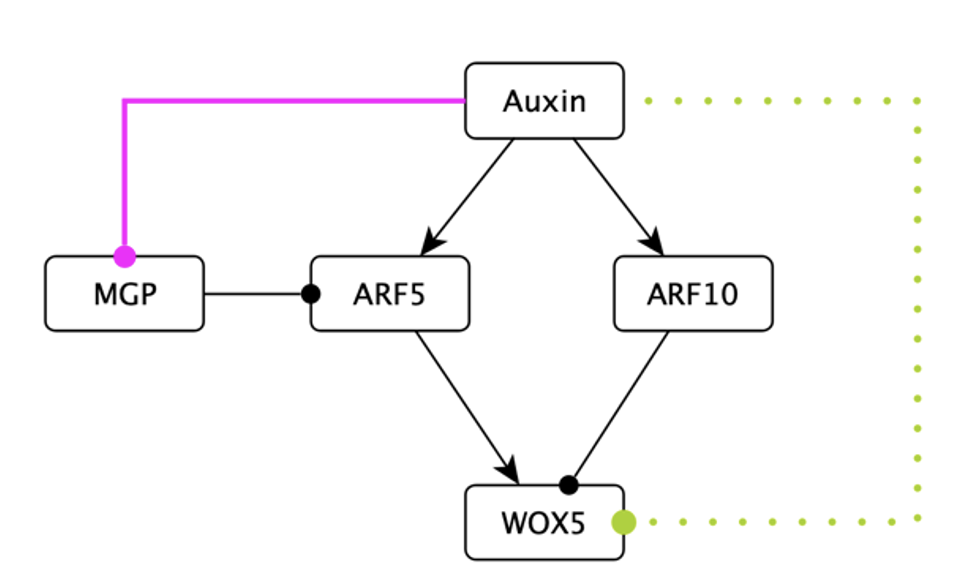
Plant root growth is driven by the activity of stem cells housed in a niche at the root tip. These stem cells self-renew and produce new cells, acting as a “growth engine”. A subset of these cells express WOX5, and maintain the surrounding cells undifferentiated. WOX5 is regulated by several factors, and network models help us understand how. Researchers have found paradoxical results regarding how auxin regulates WOX5: some experiments show that auxin promotes its expression, while others found it is repressed (Figure 1). Auxin does this opposite regulation through different ARFs (Figure 2) posing the possibility that accounting for how the ARFs themselves are regulated can reconcile the results. Here we will explore this possibility using an in silico root model.
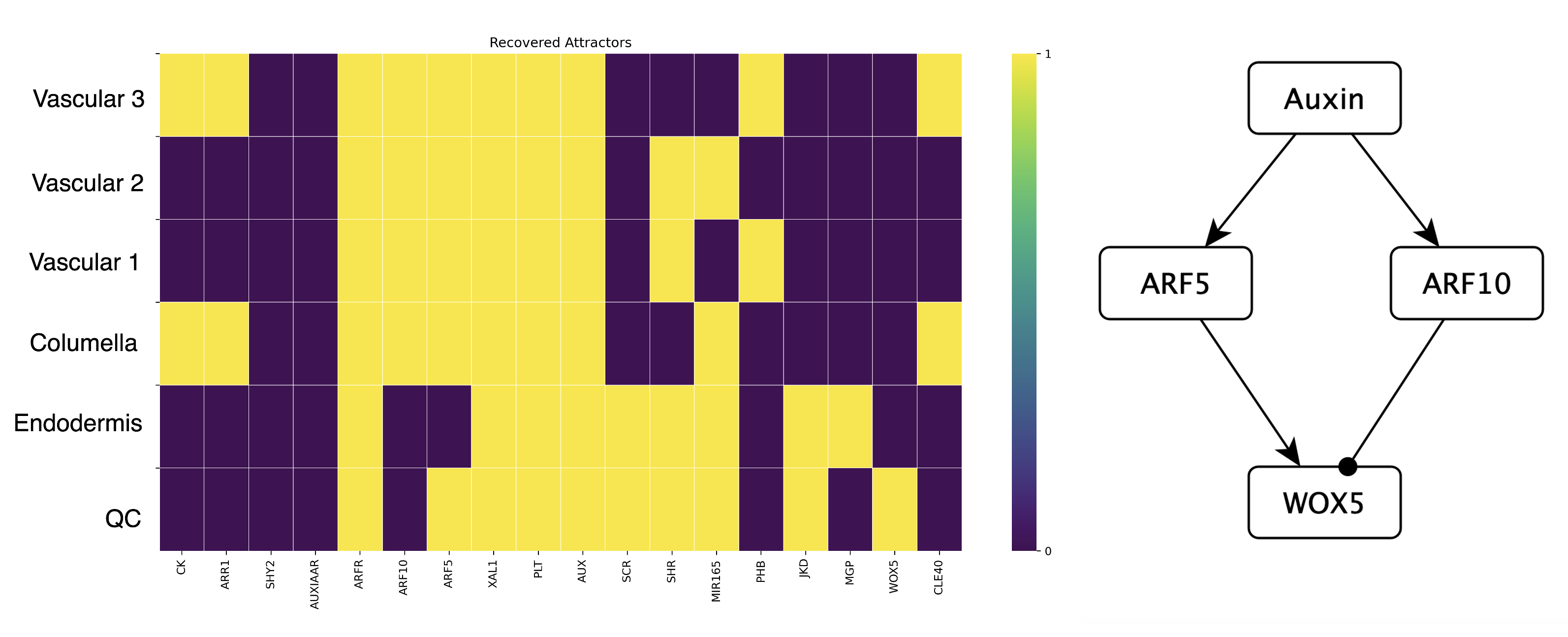
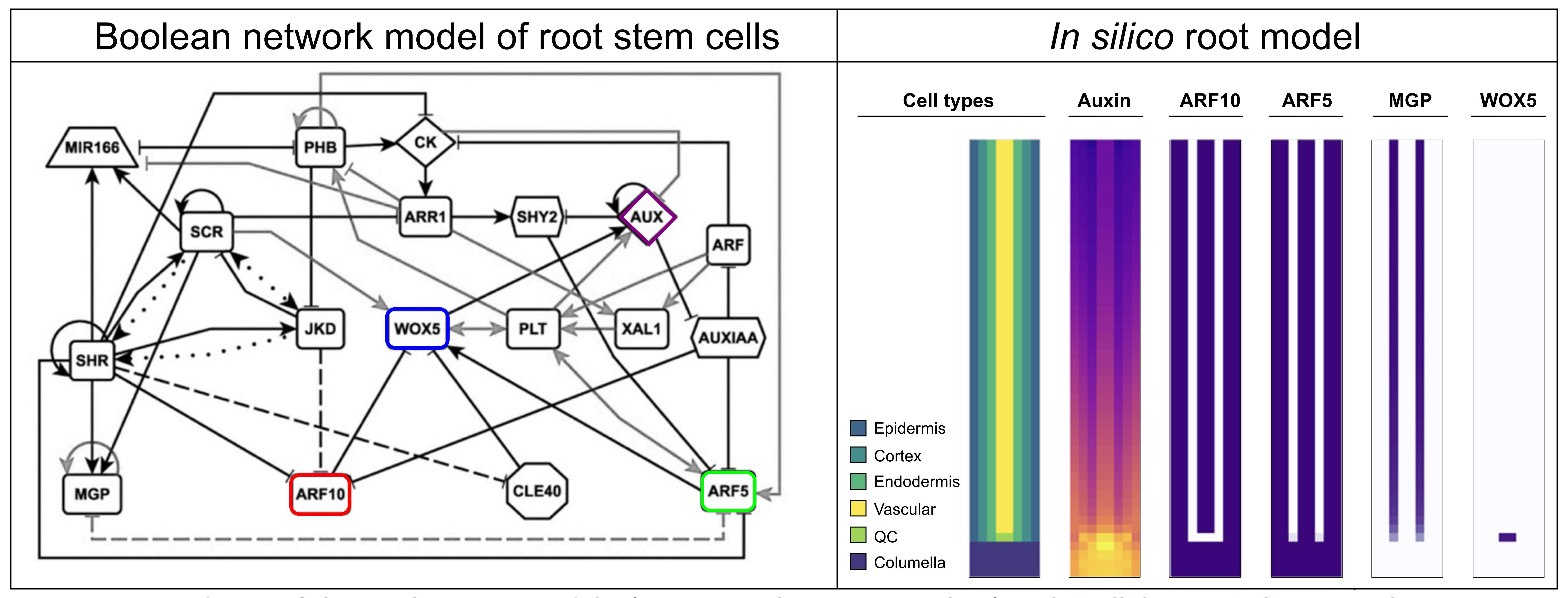
The Boolean network we used last week describes the gene and hormonal activity configurations of the cells in the root tip (Figure 3 left). This model recovers an attractor representing the columella, the QC, the endodermis, and three types of vascular cells. In this model WOX5 is regulated positively by ARF5, and negatively by ARF10, and also by CLE40. Both ARF5 and ARF10 are auxin response factors, activated by auxin. Moreover, other root transcription factors control ARF5 and ARF10 expression causing them to be expressed in specific cells of the root (see Figure 3). This regulation is included in the Boolean model we studied last week.
Here you will explore whether this is relevant to understanding the auxin–WOX5 paradox. For this you will use an in silico root model in which each cell carries a copy of the Boolean network initialized in a particular attractor, which allows us to test hypotheses and find an explanation.
Biology / conceptual thinking
Notice that the QC is the only attractor with WOX5 activity. This makes sense given that its logical rule is:
\(WOX5_{t+1} =NOT\) \(ARF10\) \(AND\) \(ARF5\) \(AND\) \(NOT\) \(CLE40\)
Notice how the other attractors have either ARF10 and/or CLE40 activity (repressors of WOX5), or lack ARF5 activity (activator of WOX5). The QC attractor is the only one with the following combination: ARF5 active, ARF10 inactive, CLE40 inactive; therefore allowing for WOX5 activity.
9.3 The model
Today you will use a multicellular model that simulates the cells of a root tip (Figure 3, right). The tissue is modeled as a grid, where the \(x\) position defines the radial direction and the \(y\) the longitudinal position (\(y=0\) being the cells at the TOP). The cells are organized in cell layers (8 layers, so \(x\) is 8), each with a gene regulatory network initialized in a different attractor. The model simulates the transport of auxin from cell to cell, where the cell type (the attractor) defines the direction of polar auxin transport. In figure 3 (right) you can see the different properties that the cells have: a cell type, auxin levels, gene expression (in the figure we see only ARF10, ARF5, MGP and WOX5). These properties are stored in individual grids: one grid for the cell type, another for auxin levels, and many others for each of the genes in the network.
Files
1.- Rootfunctions.py – File storing the functions used in Root-model-Auxin.py. For this practical you only need to modify rootNetwork. Yet, have a look at all of them and try to understand what they do (useful for Question 9.1).
The file contains the following functions:
findNeighbors- retrieves the \(x\) and \(y\) coordinates of all the neighbors of a given cell.auxinTransport- executes the passive and the polar auxin transport using the levels of auxin in each cell stored inauxinGrid. The direction at which a cell transports auxin is determined by its cell type (cellgrid).rootNetwork- stores the interactions of the root network. This file is similar to the one we used in the last practical. A key difference is that the auxin levels are an input that can modify gene expression (auxininput=parameters['auxininput']). You will modify this function in question 9.5.initialCondition- Initializes the grids of the model. We have many cells in this simulation, and we have a grid per cell property: cell type (cellgrid), auxin levels (auxinGrid), and a grid per variable of the network (e.g.WOX5grid).nodeUpdate- function to take the output of the ODE and stored it in the grids.plotGrids- this function plots 6 grids you provide as heatmaps. The labelling assumes you givecellgrid,auxingrid,arf10grid,arf5grid,mgpgrid,wox5gridas input.
2.- Root-model-Auxin.py – code to simulate the in silico root where we use the functions in a specific order. Before the main() we define the dimensions of the grid, the parameters, initialize the grid, and then inside the main() we use a for loop to run the auxin transport and network update for a defined number of time steps. In this loop, the output of the network is plotted every 10 steps.
Tip: if the simulation runs too slow, decrease this plotting frequency.
3.- auxin_grid.npy – initial auxin levels in each cell with the auxin gradient. This is used to initialize auxinGrid in Root-model-Auxin.py.
9.4 Questions
Exercise 9.1 (Algorithmic thinking) What are the algorithmic steps you would take if you want to make a model of the root in which there is transport of auxin between the cells every timestep, and the cells update their regulatory networks every 10 steps? Keep in mind that the network defines the cell type of the cells, which in turn defines the direction at which they transport auxin to their neighbors. Make a plain-language description, pseudocode, or flowchart using pen and paper.
Tip: have a look at the code in the Root-model-Auxin.py and Rootfunctions.py.
Then, run Root-model-Auxin.py.Let op! If the plotting updates too slow, you can change the frequency of plotting in line 56.
Exercise 9.2 (Biology) What happens if you update the network more often than the auxin transport? How would you implement this modification in the code?
Exercise 9.3 (Biology & algorithmic thinking) Now let’s use the model to simulate the auxin treatments from the experiments in Figure 1. To simulate an auxin treatment change the value of \(AuxinTreatment\) in Root-model-Auxin.py. Try values of 10 and 750. This increases auxin levels in all cells by the same amount. You can run one treatment at a time, or make a for loop and run the three in one go.
- How does auxin treatment affect gene expression in the root?
- Does the model output correspond to the transcriptional reporter of WOX5 shown in Figure 1?
- Do you see activation or repression as reported experimentally? (You can visualize different genes by changing the grid provided to
plotGrids(), although the labelling will be wrong.)
Notice the simulation creates an output folder with png images.
Exercise 9.4 (Biology & algorithmic/mathematical thinking) The model does not yet reproduce the experiments. Yet, we can use this model to test hypotheses and aim to recover such behaviours (Figure 1). Let’s implement the following two hypotheses:
Modify the equations for MGP and WOX5 in rootNetwork accordingly.
For MGP, retain its logical rule but multiply its production term by a negative regulatory term of auxin (\(Km = 45\)): \(\frac{45}{45+auxininput}\)
Similarly, add a negative regulation by auxin to the WOX5 equation (\(Km = 1000\)).
Run control and auxin treatments with (i) updated MGP, (ii) updated WOX5, and (iii) both updates. Use the plotting functions to plot the final timepoint.
It is also interesting to run the timecourse, by plotting every 10 steps.
Do you now see the differences reported by experimentalists?
Exercise 9.5 (Biological interpretation) You should observe that WOX5 expression gradually expands into the endodermis and disappears from QC cells when both equations are updated with the two hypotheses. This matches the experimental microscopy from Figure 1.
- Why do different parts of the root respond differently to auxin?
- Why does auxin regulation of MGP do? (Look at the network…)
- Why does induction of WOX5 occur mainly in “upper” cells and repression in the cells at the bottom?
Exercise 9.6 (Algorithmic thinking) Experimental repression of WOX5 was detected by RT-PCR at the very tip (Figure 1, right panel). Reproduce this in silico by saving the average WOX5 levels of cells at the tip (\(y\) < 40) at the end of the simulation for all three treatments. For comparison, also calculate the average for the whole root. Then plot this average WOX5 expression levels (y axis) for each auxinTreatment= 0, 10 and 750 (x axis).
Tip: WOX5Grid stores WOX5 levels, and \(y\) controls the position of the cells with \(y=0\) being the bottom of the root.
- What is the overall effect of auxin treatments in the root tip? Considering that the root is a mixture of many cell types, which cells best represent the plotted values?
Exercise 9.7 (Biology & algorithmic thinking) Now, let’s compare WOX5 regulation across tissues. Let’s use the model to do an in RT-PCR in silico analysis of only the cells in the QC position, and also the endodermis cells. This has not been done experimentally and thus you will generate a prediction (!).
Tip: cell types are stored in cellGrid.
- Compare what you see in the root simulation, and what you obtain in the simulated RT-PCR plot. Which case represents better how WOX5 is regulated in the root tip? Why?
Exercise 9.8 (Biology & algorithmic thinking) Changes in WOX5 expression depend on tissue context and auxin dosage. Simulate a range of auxin treatments from 0 to 750.
Can you now explain the auxin–WOX5 paradox? What mechanism do you propose?
Exercise 9.9 (Biology & conceptual thinking)
Perform an in silico intervention to alter WOX5 dosage responsiveness to auxin. For example, what would it take to enforce its continuous expression, or block completely its induction in the endodermis.
How would you test these model predictions experimentally?
Exercise 9.10 (Biological thinking) All models are wrong, but some are useful. The model we studied today offered us a mechanistic insight into the paradoxical auxin responses. Yet, (to make things more fun), the original experiments used different auxin analogs and other molecules which are transported very differently between cells: NAA (transported independently of PINs), IAA (PIN-transported), and NPA (blocks PINs, increasing auxin in the tip). How does this information change your conclusions? What extensions to the model would you make to account for these differences?